by Biostatsquid | Mar 20, 2023 | BiostatTOOLS
If you are looking for tools to carry out differential gene expression analysis, check out my top gene expression analysis tools and bioinformatic resources. If you are new here, don’t forget to check out Biostatsquid’s guided tutorials and step-by-step...

by Biostatsquid | Mar 18, 2023 | BiostatTOOLS
If you are just starting to code in R or Python or would like to improve your coding skills, check out my top coding resources and tools for beginners (and not so beginners!). Of course, here at Biostatsquid you can also find guided tutorials and step-by-step guides...
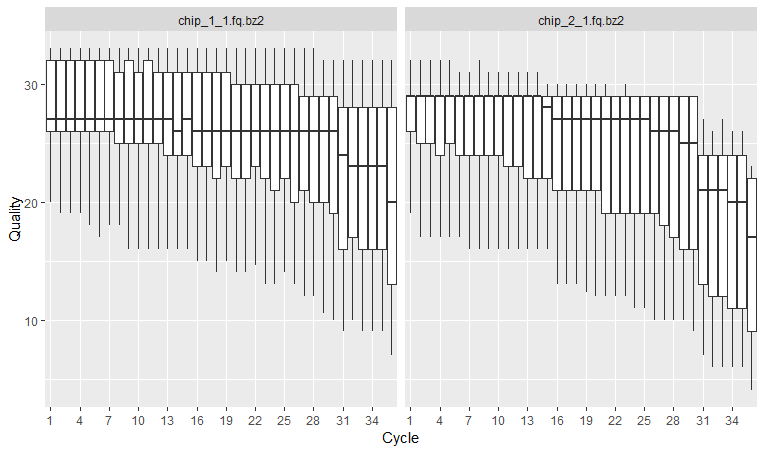
by Biostatsquid | Feb 8, 2023 | Tutorials
R TUTORIAL: From .fastq to .bam, preprocess and align your sequencing reads with this step-by-step sequencing reads R tutorial Table of contents 5 Before you start 5 Sequencing reads processing pipeline: complete script 5 Step-by-step guide for sequencing read...
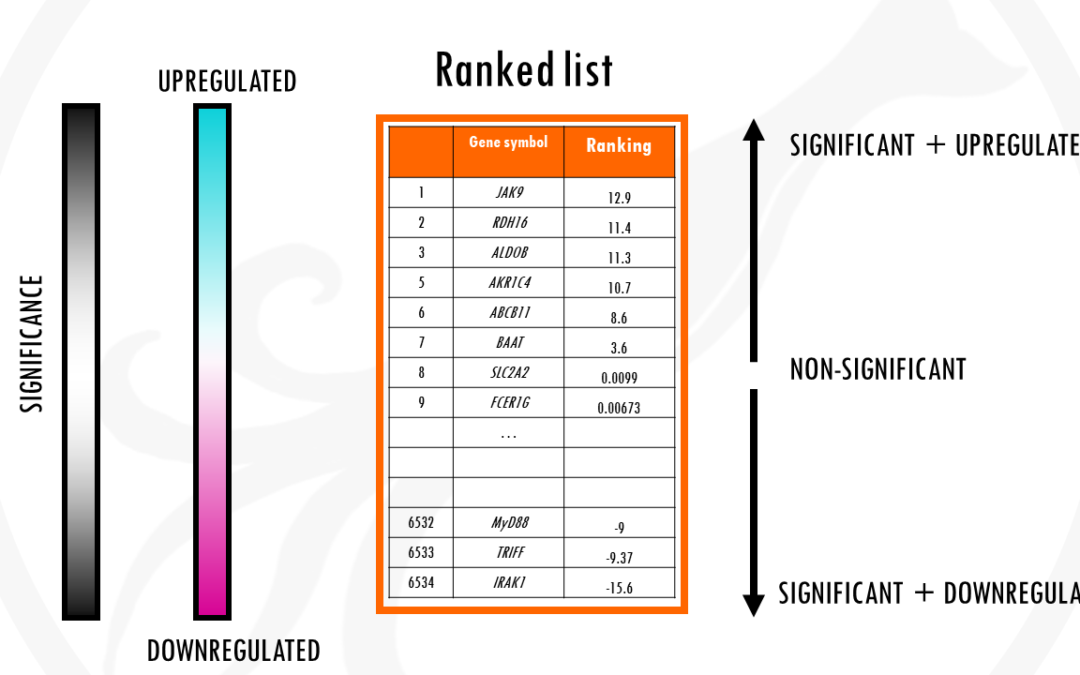
by Biostatsquid | Jan 23, 2023 | Learning, RNAseq, scRNAseq, Statistics
What is gene set enrichment analysis and how can you use it to summarise your differential gene expression analysis results? This post will give you a simple and practical explanation of Gene Set Enrichment Analysis, or GSEA for short. You will find out: What is Gene...
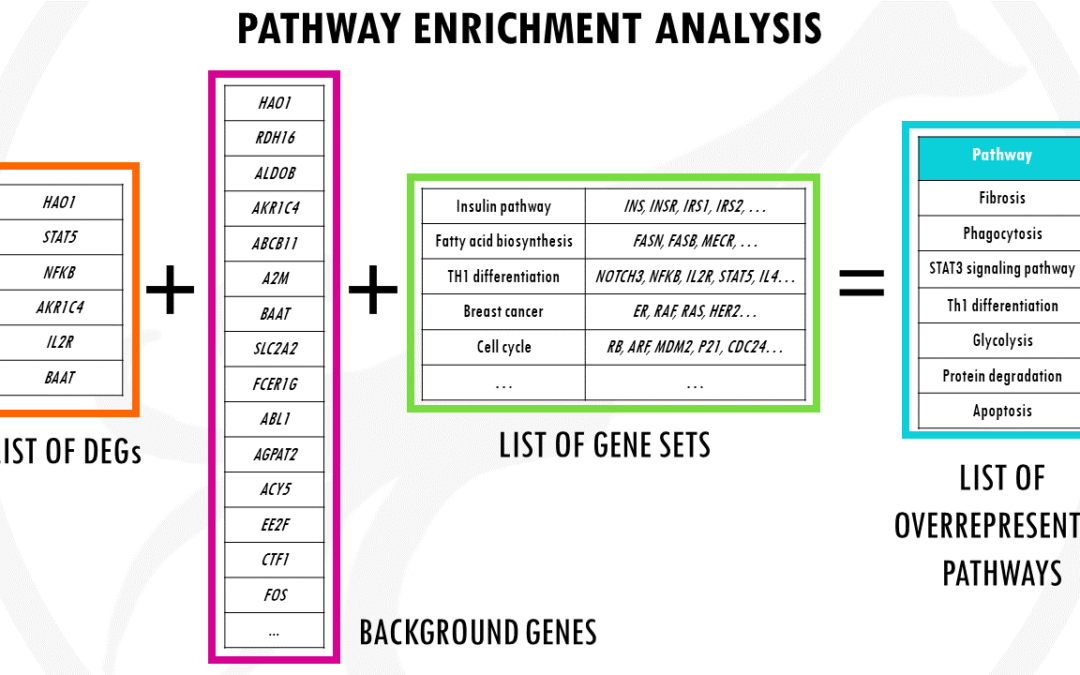
by Biostatsquid | Jan 23, 2023 | Learning, RNAseq, scRNAseq, Statistics
An overview of pathway enrichment analysis and how you can use it for your differential gene expression analysis data. In this post, you will find pathway enrichment analysis explained in a simple way with examples. I will try to give you a simple and practical...







