by Biostatsquid | Apr 9, 2026 | Learning, Machine learning, Statistics
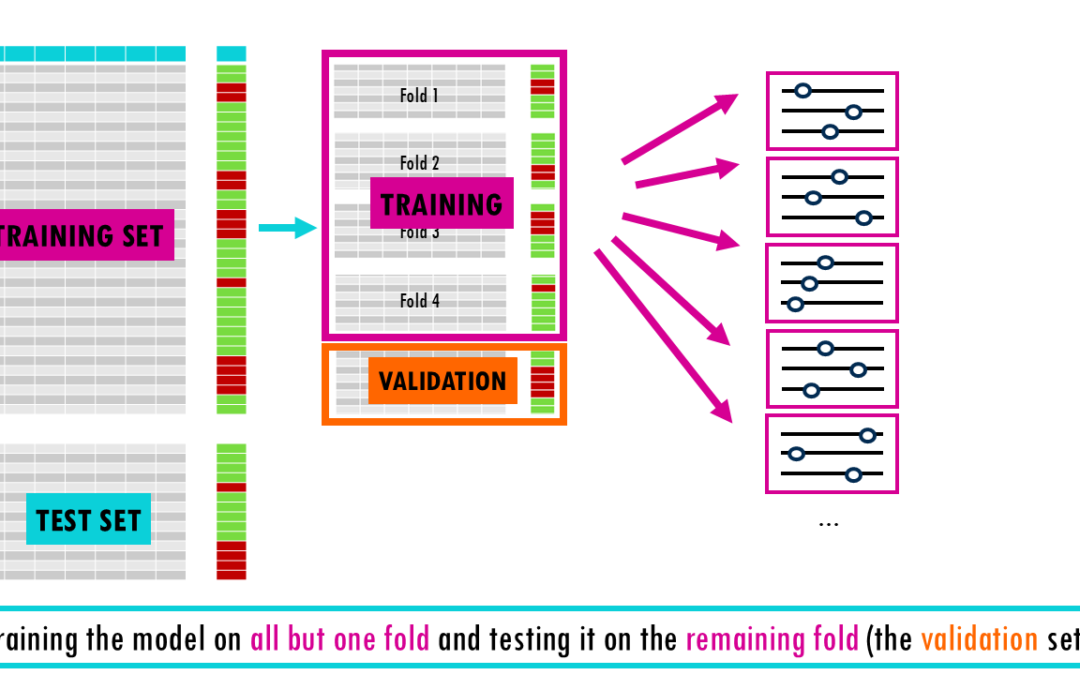
From a simple train-test split to stratified nested cross-validation! Imagine studying for a final exam by memorizing the exact answers to a practice test. You might feel like a genius while grading yourself, but if the actual exam asks the same questions in a...
by Biostatsquid | Apr 1, 2026 | Learning, scRNAseq
One of the most common questions in single-cell RNA sequencing analysis is deceptively straightforward: how many cells express a given gene, and are the expression levels truly comparable across datasets? If you have ever stared at your results wondering whether your...
by Biostatsquid | Feb 21, 2026 | Learning, Statistics
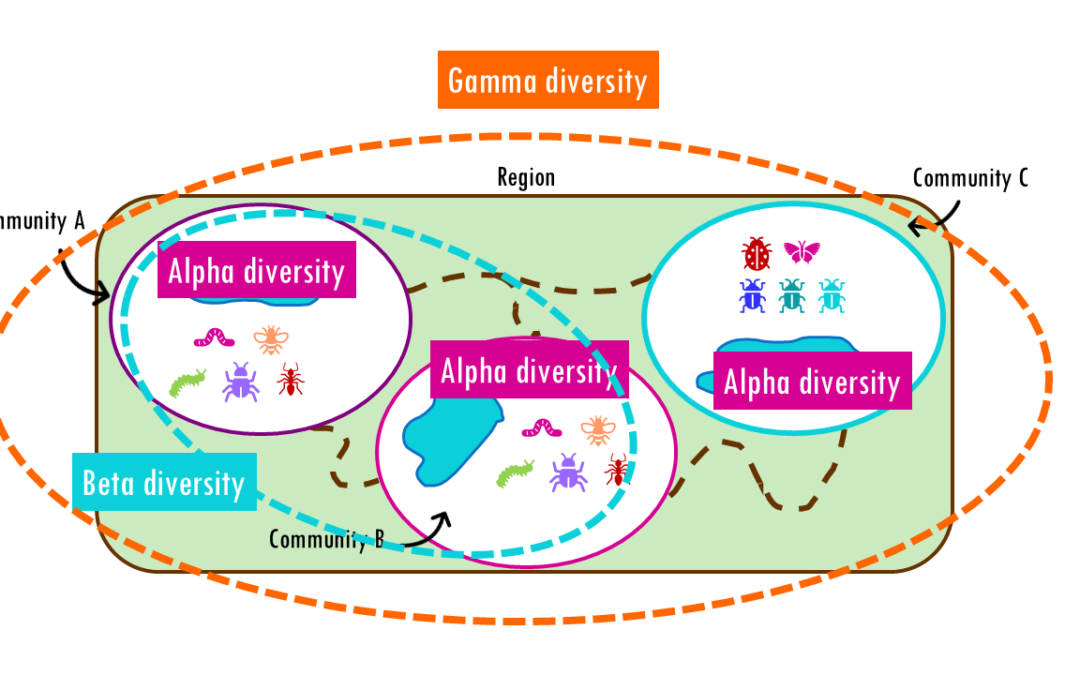
How different are communities from each other? Beta diversity easily explained! If you’ve ever hiked from a dense forest into an open grassland, you’ve probably noticed how dramatically the plants, insects, and animals can change within just a few miles. This...
by Biostatsquid | Feb 14, 2026 | Learning, Statistics
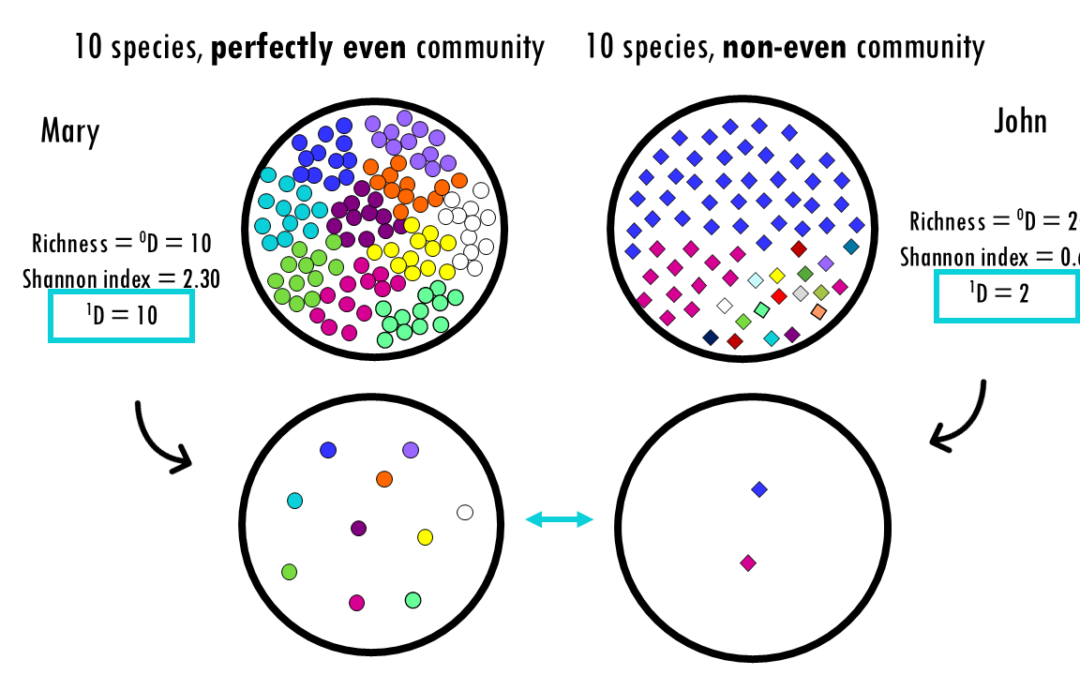
Exploring Hill numbers to compare the diversity across communities Whether you are analyzing a rainforest ecosystem, a human gut microbiome, or a B-cell receptor (BCR) repertoire, the fundamental challenge is the same: How do we accurately measure the diversity within...
by Biostatsquid | Feb 14, 2026 | Learning, Statistics
Exploring alpha diversity indices and how to interpret them: Shannon, Simpson, Gini, Chao1 and more! Whether you are analyzing a rainforest ecosystem, a human gut microbiome, or a B-cell receptor (BCR) repertoire, the fundamental challenge is the same: How do we...
by Biostatsquid | Dec 14, 2025 | Learning, scRNAseq
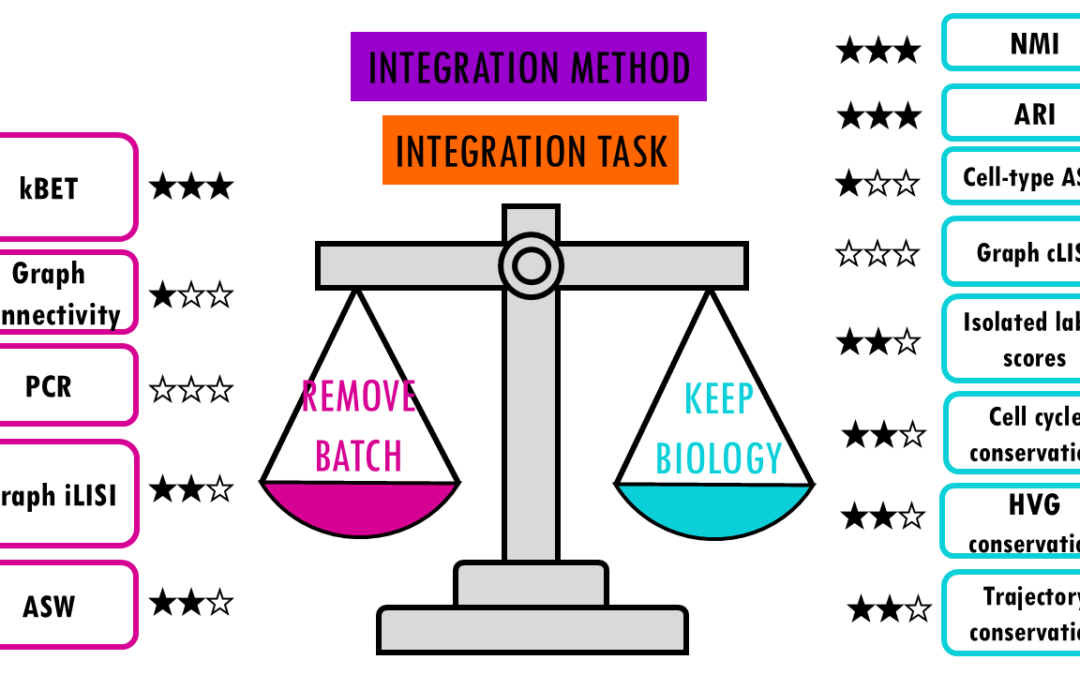
Comparing top integration methods for scRNAseq data When we want to combine multiple scRNA-seq datasets to answer bigger questions, we encounter batch effects – unwanted technical variations that arise from differences in how the experiments were performed....






