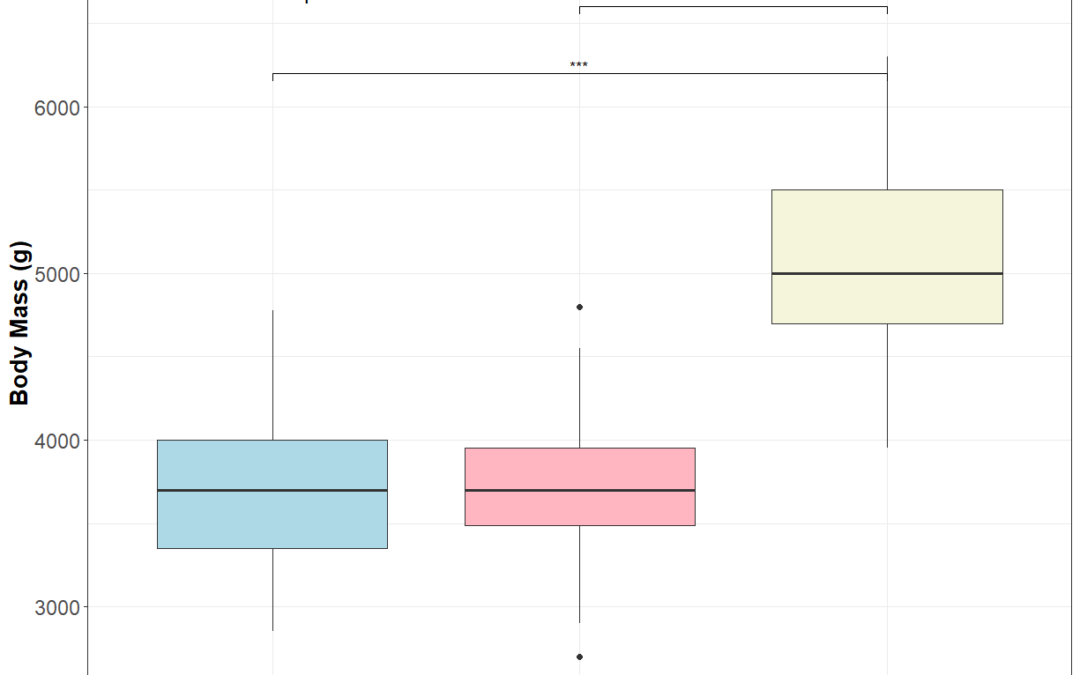
by Biostatsquid | Jun 6, 2025 | Learning, Statistics, Statistics in R, Tutorials
When working with biological data, we often want to compare measurements across multiple groups. However, these measurements aren’t always normally distributed. In such cases, non-parametric methods like the Kruskal-Wallis test and Dunn’s post-hoc test are ideal...

by Biostatsquid | May 5, 2025 | scRNAseq, Tutorials
A step-by-step easy R tutorial to preprocess scRNAseq data with Seurat v5 In this easy, step-by-step tutorial you will learn how a Seurat object is structured and how to preprocess scRNAseq data using the standard workflow with Seurat v5. This is a hands-on...

by Biostatsquid | Apr 27, 2025 | Learning, RNAseq, scRNAseq
Understanding the structure of Seurat objects version 5 – step-by-step simple explanation! If you’ve worked with single-cell RNAseq data, you’ve probably heard about Seurat. In this blogpost, we’ll cover the the Seurat object structure,in...
by Biostatsquid | Apr 24, 2025 | scRNAseq, Statistics
SCTransform (Single-Cell Transform) is a normalization method primarily used in scRNA-seq data analysis. It was developed to address limitations in standard normalization approaches when dealing with single-cell data. You can check how to apply SCTransform on your...
by Biostatsquid | Apr 23, 2025 | Learning, Statistics
That’s a really good and very common question in differential gene expression analysis! It feels intuitive that the larger the difference in expression (log fold change, or logFC), the more significant it should be (i.e., the smaller the p-value), but that’s not...





